One of the best explanations of how modern evolutionary genomics is done is in the fourth chapter of Richard Dawkins's fantastic The Ancestor's Tale. The book starts with humans (although he makes the point that he could have started with any other species on Earth), and tracks backwards in time to each of the points where the human lineage intersects with other lineages. So it starts out with chapters about our nearest relatives -- bonobos and chimps -- and gradually progresses to more and more distantly-related groups, until by the last chapter we've united our lineage with every other life form on the planet.
In chapter four ("Gibbons"), he describes something of the methodology of how this is done, using as an analogy how linguists have traced the "ancestry" (so to speak) of the surviving copies of Chaucer's The Canterbury Tales, each of which have slight variations from the others. The question he asks is how we could tell what the original version looked like; put another way, which of those variations represent alterations, and which were present in the first edition.
The whole thing is incredibly well done, in the lucid style for which Dawkins has rightly become famous, and I won't steal his thunder by trying to recap it here (in fact, you should simply read the book, which is wonderful from beginning to end). But a highly oversimplified capsule explanation is that the method relies on the law of parsimony -- that the model which requires the fewest ad hoc assumptions is the most likely to be correct. When comparing pieces of DNA from groups of related species, the differences come from mutations; but if two species have different base pairs at a particular position, which was the original and which the mutated version -- or are both mutations from a third, different, base pair at that position?
The process takes the sequences and puts together various possible "family trees" for the DNA; the law of parsimony states that the likeliest one is the arrangement that requires the fewest de novo mutations. To take a deliberately facile example, suppose that within a group of twelve related species, in a particular stretch of DNA, eleven of them have an A/T pair at the third position, and the twelfth has a C/G pair. Which is more likely -- that the A/T was the base pair in the ancestral species and species #12 had a mutation to C/G, or that C/G was the base pair in the ancestral species and species #1-11 all independently had mutations to A/T?
Clearly the former is (hugely) more likely. Most situations, of course, aren't that clear-cut, and there are complications I won't go into here, but that's the general idea. Using software -- none of this is done by hand any more -- the most parsimonious arrangement is identified, and in the absence of any evidence to the contrary, is assumed to be the lineage of the species in question.
This is pretty much how all cladistics is done. Except in cases where we don't have DNA evidence -- such as with prehistoric animals known only from fossils -- evolutionary biologists don't rely much on structure any longer. As Dawkins himself put it, "Even if we were to erase every fossil from the Earth, the evidence for evolution from genetics alone would be overwhelming."
The reason this comes up is a wonderful study that came out this week in Science that uses these same techniques to put together the ancestry of all the modern varieties of grapes. A huge team at the Karlsruher Institut für Technologie and the Chinese Yunnan Agricultural University analyzed the genomes of 3,500 different grapevines, including both wild and cultivated varieties, and was able to track their ancestry back to the southern Caucasus in around 11,000 B.C.E. (meaning that grapes seem to have been cultivated before wheat was). From there, the vine rootstocks were carried both ways along the Silk Road, spreading all the way from China to western Europe in the process.
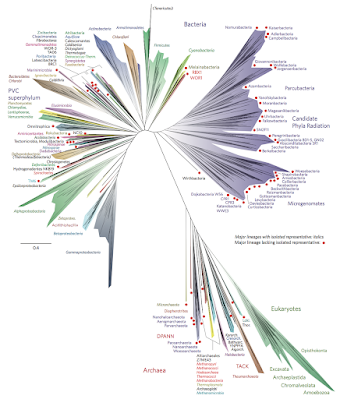
There are a lot of things about this study that are fascinating. First, of course, is that we can use the current assortment of wild and cultivated grape vines to reconstruct a family tree that goes back thirteen thousand years -- and come up with a good guess about where the common ancestor of all of them lived. Second, though, is the more general astonishment at how sophisticated our ability to analyze genomes has become. Modern genomic analysis has allowed us to create family trees of all living things that boggle the mind -- like this one:
These sorts of analyses have overturned a lot of our preconceived notions about our place in the world. It upset a good many people, for some reason, when it was found we have a 98.7% overlap in our DNA with our nearest relatives (bonobos) -- that remaining 1.3% accounts for the entire genetic difference between yourself and a bonobo. People were so used to believing there was a qualitative biological difference between humans and everything other living thing that to find out we're so closely related to apes was a significant shock. (It still hasn't sunk in for some people; you'll still hear the phrase "human and animal" used, as if we weren't ourselves animals.)
Anyhow, an elegant piece of research on the ancestry of grapes is what got all this started, and after all of my circumlocution you probably feel like you need a glass of wine. Enjoy -- in vino veritas, as the Romans put it, even if they may not have known as much about where their vino originated as we do.
****************************************


No comments:
Post a Comment